Please insert your variant position in hg38-coordinates (1-based); use just a number/letter without a leading "chr" for the chromosome (e.g. "1" or "X" would be valid).
| CHROM(hg38) | POS(1-based, hg38) | ref | alt | aaref | aaalt | rs_dbSNP | CHROM(hg19) | POS(1-based, hg19) | CHROM:POS:REF:ALT(hg38) | genename(s) | Uniprot_ID mapped to AlphaFold2 | Uniprot_IDs | HGVSp_VEP mapped to AlphaFold2 | HGVSp_VEPs | CADD_raw (this is not CADD_phred) | REVEL score | DEOGEN2 scores (values separated by semicolon for the Uniprot-IDs/Genes) | AlphaFold2's pLDDT | SOLVENT_ACCESSIBILITY_core | in_gnomad_train (training dataset, TRUE if yes) | in_clinvar_ds (mostly testing dataset, TRUE if yes) | AlphScore_final | AlphScore_final+CADD | AlphScore_final+REVEL | REVEL+CADD | AlphScore_final+REVEL+CADD | AlphScore_final+DEOGEN2 | CADD+DEOGEN2 | DEOGEN2+REVEL | AlphScore_final+DEOGEN2+REVEL | AlphScore_final+CADD+DEOGEN2 | CADD+DEOGEN2+REVEL |
|---|
| Column | Column description |
|---|---|
| CHROM(hg38) | chromosome (hg38) |
| POS(1-based, hg38) | position (hg38) |
| ref | reference allele |
| alt | alternative allele |
| aaref | reference amino acid |
| aaalt | alternative amino acid |
| rs_dbSNP | rs number |
| CHROM(hg19) | chromosome (hg19) |
| POS(1-based, hg19) | position (hg19) |
| CHROM:POS:REF:ALT(hg38) | variant id in the format: chromosome:position:reference amino acid:alternative amino acid |
| genename(s) | genename, taken from dbNSFP |
| Uniprot_ID mapped to AlphaFold2 | The Uniprot-IDs of the structural models that were used to create AlphScore_final (multiple entries separated by ; ) |
| Uniprot_IDs | Uniprot_acc as provided by dbNSFP |
| HGVSp_VEP mapped to AlphaFold2 | The missense variant(s) as used to create AlphScore_final; these variant(s) correspond(s) to the Uniprot_acc_split |
| HGVSp_VEPs | HGVSp_VEP as provided by dbNSFP |
| CADD_raw (this is not CADD_phred) | CADD_raw as provided by dbNSFP |
| REVEL score | REVEL_score as provided by dbNSFP |
| DEOGEN2 scores (values separated by semicolon for the Uniprot-IDs/Genes) | DEOGEN2_score as provided by dbNSFP |
| AlphaFold2's pLDDT | AlphaFold's pLDDT-score of the residue (if a variant affects multiple proteins, the values of the proteins as indicated in Uniprot_acc_split are given separated by ; ). |
| SOLVENT_ACCESSIBILITY_core | Solvent accessibility of the residue as calculated for C-alpha by DSSP (if a variant affects multiple proteins, the values of the proteins as indicated in Uniprot_acc_split are given separated by ; ). |
| in_gnomad_train (training dataset, TRUE if yes) | TRUE if the variant was in the gnomAD set used for training |
| in_clinvar_ds (mostly testing dataset, TRUE if yes) | TRUE if the variant was in the ClinVar set used for validation / training of combined scores |
| AlphScore_final | This column corresponds to AlphScore_final |
| AlphScore_final+CADD | This column corresponds to AlphScore_final + CADD |
| AlphScore_final+REVEL | This column corresponds to AlphScore_final + REVEL |
| REVEL+CADD | This column corresponds to REVEL + CADD |
| AlphScore_final+REVEL+CADD | This column corresponds to AlphScore_final + REVEL + CADD |
| AlphScore_final+DEOGEN2 | This column corresponds to AlphScore_final + DEOGEN2 |
| CADD+DEOGEN2 | This column corresponds to CADD + DEOGEN2 |
| DEOGEN2+REVEL | This column corresponds to DEOGEN2 + REVEL |
| AlphScore_final+DEOGEN2+REVEL | This column corresponds to AlphScore_final + DEOGEN2 + REVEL |
| AlphScore_final+CADD+DEOGEN2 | This column corresponds to AlphScore_final + CADD + DEOGEN2 |
| CADD+DEOGEN2+REVEL | This column corresponds to CADD + DEOGEN2 + REVEL |
A graphical summary for the individual scores and combinations of up to two scores is given by the following density plot (Figure S10 from the manuscript):
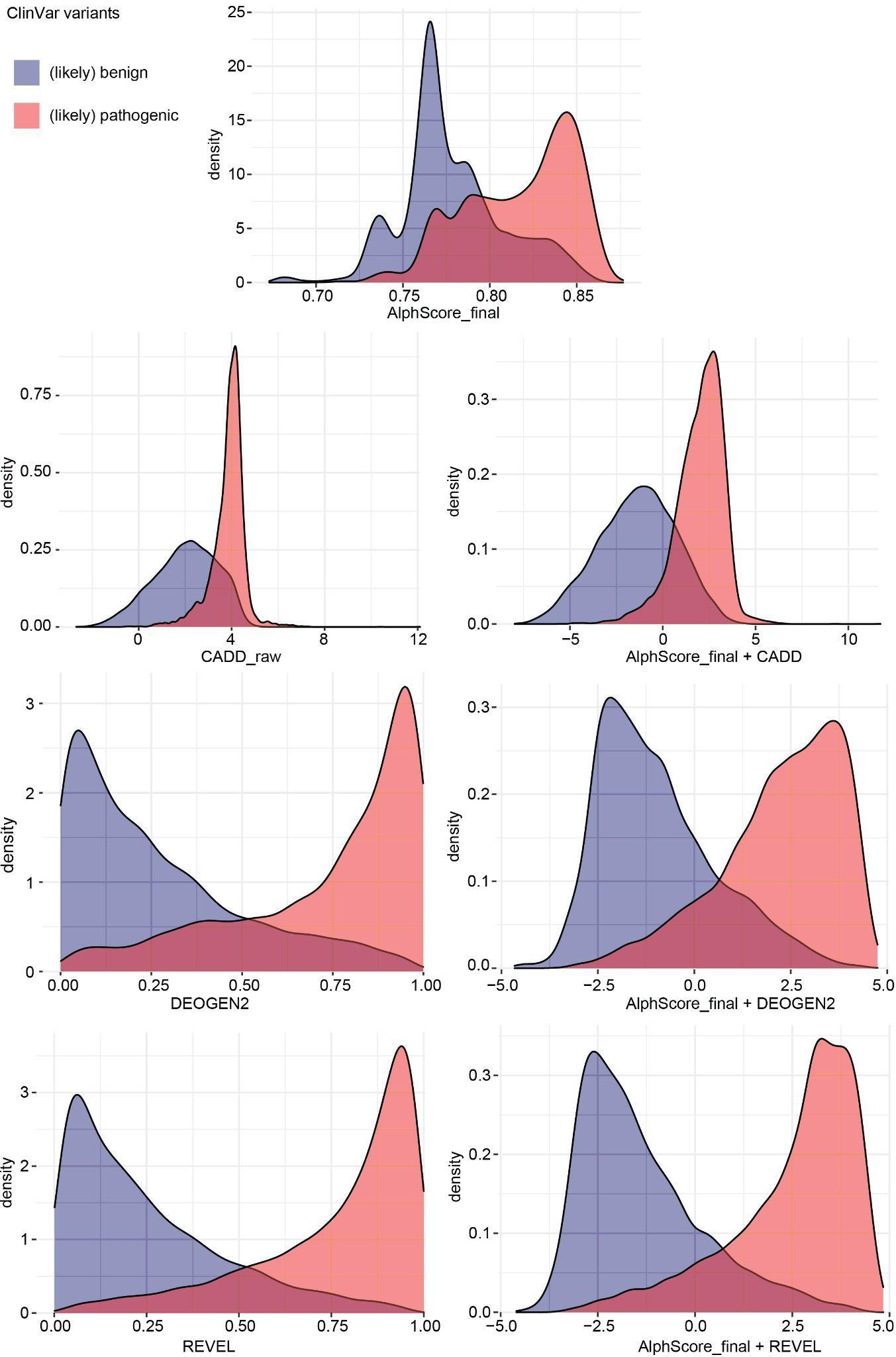
See Supplementary Table S6 for cut-offs and performance metrices of each score.
The data that can be accessed via this site is taken from zenodo (https://doi.org/10.5281/zenodo.6288138) and is described in:
Axel Schmidt, Sebastian Röner, Karola Mai, Hannah Klinkhammer, Martin Kircher, Kerstin U Ludwig, Predicting the pathogenicity of missense variants using features derived from AlphaFold2, Bioinformatics, 2023;, btad280, https://doi.org/10.1093/bioinformatics/btad280